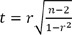|
 |
- Search
| Korean J Fam Med > Volume 35(5); 2014 > Article |
In this section, we explain the assumptions in the analysis of covariance and the relationship between the magnitudes of correlation coefficient and P-value, which appeared in the articles titled, "Association between nutrition label reading and nutrient Intake in Korean adults: Korea National Health and Nutritional Examination Survey, 2007-2009 (KNHANES IV)," by Kim et al.1) and "Association between appendicular fat mass and metabolic risk factors," by Park et al.2) published in July 2014.
Analysis of covariance (ANCOVA) is a general linear model which blends ANOVA and regression.3) We use ANCOVA to evaluate whether means of a outcome variable are equal across levels of a categorical independent variable (so-called groups), while controlling for the effects of other continuous variables that are not of primary interest, known as covariates. Therefore, when performing ANCOVA, the means of outcome variable are adjusted to what they would be if all groups were equal on the covariates (least square means).
There are three important assumptions that underlie the use of ANCOVA. 1) The residuals (error terms) should be normally distributed. 2) The error variances should be equal for all groups. 3) The slopes of the different regression lines should be equivalent, i.e., regression lines should be parallel among groups.
The third assumption, concerning the homogeneity of different treatment regression slopes is particularly important in evaluating the appropriateness of ANCOVA model. This assumption also implies that all covariates should be confounding variables, i.e., there are no interactions between group and covariates.
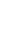
Pearson's correlation coefficient is defined as the covariance of the two variables divided by the product of their standard deviations and a measure of the strength of linear relationship between two normally distributed variables. For two variables from an uncorrelated bivariate normal distribution, the sampling distribution of Pearson's correlation coefficient follows Student t-distribution with degrees of freedom n - 2. Specifically, if the underlying variables have a bivariate normal distribution, the test statistic has a Student t-distribution under the null hypothesis (zero correlation). This also holds approximately even if the observed values are non-normal, provided sample sizes are not very small. Thus this test statistic could be used for Spearman's rank correlation.
As we can see from above formula, test statistic of correlation coefficient is the function of a sample size (n) and a correlation coefficient. Thus, under the condition of the same n, values of test statistics are proportional to those of correlation coefficients and the absolute values of correlation coefficient should be inversely proportional to those of P-value.
References
1. Kim MG, Oh SW, Han NR, Song DJ, Um JY, Bae SH, et al. Association between nutrition label reading and nutrient intake in Korean adults: Korea National Health and Nutritional Examination Survey, 2007-2009 (KNHANES IV). Korean J Fam Med 2014;35:190-198. PMID: 25120890.



2. Park SY, Kwon KY, Kim JH, Choi HH, Han KH, Han JH. Association between appendicular fat mass and metabolic risk factors. Korean J Fam Med 2014;35:182-189. PMID: 25120889.



3. Snedecor GW, Cochran WG. Statistical methods. 2nd ed. Ames: Iowa State University Press; 1980.
- TOOLS
-
METRICS

-
- 0 Crossref
- Scopus
- 3,578 View
- 20 Download
- Related articles in KJFM
-
Comments on Statistical Issues in January 20162016 January;37(1)
Comments on Statistical Issues in November 20152015 November;36(6)
Comments on Statistical Issues in January 20152015 January;36(1)
Comments on Statistical Issues in September 20152015 September;36(5)